mmfutils API
Contents:
MMF Utils
Small set of utilities: containers and interfaces.
This package provides some utilities that I tend to rely on during development. Presently it includes some convenience containers, plotting tools, and a patch for including zope.interface documentation in a notebook.
(Note: If this file does not render properly, try viewing it through nbviewer.org)
Documentation: http://mmfutils.readthedocs.org
Source:
https://alum.mit.edu/www/mforbes/hg/forbes-group/mmfutils: Permalink (will forward).
https://gitlab.com/ColdAtoms/utilities/mmfutils: Current, in case the permalink fails.
https://hg.iscimath.org/forbes-group/mmfutils: Old, in case the permalink fails.
https://github.com/forbes-group/mmfutils: Public read-only mirror.
Issues: https://alum.mit.edu/www/mforbes/hg/forbes-group/mmfutils/issues
Build Status
Installing
This package can be installed from PyPI:
python3 -m pip install mmfutils
python3 -m pip install mmfutils[fftw] # If you have the FFTW libraries installed
or, if you need to install from source, you can get it from one of the repositories:
python3 -m pip install hg+https://alum.mit.edu/www/mforbes/hg/forbes-group/mmfutils
python3 -m pip install git+https://gitlab.com/coldatoms/utilities/mmfutils
Usage
Containers
ObjectBase and Object
The ObjectBase and Object classes provide some useful features
described below. Consider a problem where a class is defined through a
few parameters, but requires extensive initialization before it can be
properly used. An example is a numerical simulation where one passes the
number of grid points \(N\) and a length \(L\), but the
initialization must generate large grids for efficient use later on.
These grids should be generated before computations begin, but should
not be re-generated every time needed. They also should not be pickled
when saved to disk.
Deferred initialization via the ``init()`` method: The idea here
changes the semantics of __init__() slightly by deferring any
expensive initialization to init(). Under this scheme,
__init__() should only set and check what we call picklable
attributes: these are parameters that define the object (they will be
pickled in Object below) and will be stored in a list
self.picklable_attributes which is computed at the end of
ObjectBase.__init__() as the list of all keys in __dict__. Then,
ObjectBase.__init__() will call init() where all remaining
attributes should be calculated.
This allows users to change various attributes, then reinitialize the
object once with an explicit call to init() before performing
expensive computations. This is an alternative to providing complete
properties (getters and setters) for objects that need to trigger
computation. The use of setters is safer, but requires more work on the
side of the developer and can lead to complex code when different
properties depend on each other. The approach here puts all computations
in a single place. Of course, the user must remember to call init()
before working with the object.
To facilitate this, we provide a mild check in the form of an
initialized flag that is set to True at the end of the base
init() chain, and set to False if any variables are in
pickleable_attributes are set.
Serialization and Deferred Initialization: The base class
ObjectBase does not provide any pickling services but does provide a
nice representation. Additional functionality is provided by Object
which uses the features of ObjectBase to define __getstate__()
and __setstate__() methods for pickling which pickle only the
picklable_attributes. Note: unpickling an object will not call
__init__() but will call init() giving objects a chance to
restore the computed attributes from pickles.
Note: Before using, consider if these features are really needed – with all such added functionality comes additional potential failure modes from side-interactions. The ``ObjectBase`` class is quite simple, and therefore quite safe, while ``Object`` adds additional functionality with potential side-effects. For example, a side-effect of support for pickles is that ``copy.copy()`` will also invoke ``init()`` when copying might instead be much faster. Thus, we recommend only using ``ObjectBase`` for efficient code.
Object Example
ROOTDIR = !hg root
ROOTDIR = ROOTDIR[0]
import sys
sys.path.insert(0, ROOTDIR)
import numpy as np
from mmfutils.containers import ObjectBase, ObjectMixin
class State(ObjectBase):
_quiet = False
def __init__(self, N, L=1.0, **kw):
"""Set all of the picklable parameters, in this case, N and L."""
self.N = N
self.L = L
# Now register these and call init()
super().__init__(**kw)
if not self._quiet:
print("__init__() called")
def init(self):
"""All additional initializations"""
if not self._quiet:
print("init() called")
dx = self.L / self.N
self.x = np.arange(self.N, dtype=float) * dx - self.L / 2.0
self.k = 2 * np.pi * np.fft.fftfreq(self.N, dx)
# Set highest momentum to zero if N is even to
# avoid rapid oscillations
if self.N % 2 == 0:
self.k[self.N // 2] = 0.0
# Calls base class which sets self.initialized
super().init()
def compute_derivative(self, f):
"""Return the derivative of f."""
return np.fft.ifft(self.k * 1j * np.fft.fft(f)).real
s = State(256)
print(s) # No default value for L
init() called
__init__() called
State(L=1.0, N=256)
s.L = 2.0
print(s)
State(L=2.0, N=256)
One feature is that a nice repr() of the object is produced. Now
let’s do a calculation:
f = np.exp(3 * np.cos(2 * np.pi * s.x / s.L)) / 15
df = (
-2.0
* np.pi
/ 5.0
* np.exp(3 * np.cos(2 * np.pi * s.x / s.L))
* np.sin(2 * np.pi * s.x / s.L)
/ s.L
)
np.allclose(s.compute_derivative(f), df)
False
Oops! We forgot to reinitialize the object… (The formula is correct, but the lattice is no longer commensurate so the FFT derivative has huge errors).
print(s.initialized)
s.init()
assert s.initialized
f = np.exp(3 * np.cos(2 * np.pi * s.x / s.L)) / 15
df = (
-2.0
* np.pi
/ 5.0
* np.exp(3 * np.cos(2 * np.pi * s.x / s.L))
* np.sin(2 * np.pi * s.x / s.L)
/ s.L
)
np.allclose(s.compute_derivative(f), df)
False
init() called
True
Here we demonstrate pickling. Note that using Object makes the
pickles very small, and when unpickled, init() is called to
re-establish s.x and s.k. Generally one would inherit from
Object, but since we already have a class, we can provide pickling
functionality with ObjectMixin:
class State1(ObjectMixin, State):
pass
s = State(N=256, _quiet=True)
s1 = State1(N=256, _quiet=True)
import pickle, copy
s_repr = pickle.dumps(s)
s1_repr = pickle.dumps(s1)
print(f"ObjectBase pickle: {len(s_repr)} bytes")
print(f"ObjectMixin pickle: {len(s1_repr)} bytes")
ObjectBase pickle: 4397 bytes
ObjectMixin pickle: 103 bytes
Note, however, that the speed of copying is significantly impacted:
%timeit copy.copy(s)
%timeit copy.copy(s1)
1.28 μs ± 11.8 ns per loop (mean ± std. dev. of 7 runs, 1,000,000 loops each)
10.6 μs ± 9.35 ns per loop (mean ± std. dev. of 7 runs, 100,000 loops each)
Another use case applies when init() is expensive. If \(x\) and
\(k\) were computed in __init__(), then using properties to
change both \(N\) and \(L\) would trigger two updates. Here we
do the updates, then call init(). Good practice is to call
init() automatically before any serious calculation to ensure that
the object is brought up to date before the computation.
s.N = 64
s.L = 2.0
s.init()
Finally, we demonstrate that Object instances can be archived using
the persist package:
import persist.archive
a = persist.archive.Archive(check_on_insert=True)
a.insert(s=s)
d = {}
exec(str(a), d)
d["s"]
State(L=2.0, N=64, _quiet=True)
Container
The Container object is a slight extension of Object that
provides a simple container for storing data with attribute and
iterative access. These implement some of the Collections Abstract Base
Classes from the python standard
library.
The following containers are provided:
Container: Bare-bones container extending theSized,Iterable, andContainerabstract ase classes (ABCs) from the standardcontainerslibrary.ContainerList: Extension that acts like a tuple/list satisfying theSequenceABC from thecontainerslibrary (but not theMutableSequenceABC. Although we allow setting and deleting items, we do not provide a way for insertion, which breaks this interface.)ContainerDict: Extension that acts like a dict satisfying theMutableMappingABC from thecontainerslibrary.
These were designed with the following use cases in mind:
Returning data from a function associating names with each data. The resulting
ContainerListwill act like a tuple, but will support attribute access. Note that the order will be lexicographic. One could use a dictionary, but attribute access with tab completion is much nicer in an interactive session. Thecontainers.nametuplegenerator could also be used, but this is somewhat more complicated (though might be faster). Also, named tuples are immutable - here we provide a mutable object that is picklable etc. The choice betweenContainerListandContainerDictwill depend on subsequent usage. Containers can be converted from one type to another.
Container Examples
from mmfutils.containers import Container
c = Container(a=1, c=2, b="Hi there")
print(c)
print(tuple(c))
Container(a=1, b='Hi there', c=2)
(1, 'Hi there', 2)
# Attributes are mutable
c.b = "Ho there"
print(c)
Container(a=1, b='Ho there', c=2)
# Other attributes can be used for temporary storage but will not be pickled.
import numpy as np
c.large_temporary_array = np.ones((256, 256))
print(c)
print(c.large_temporary_array)
Container(a=1, b='Ho there', c=2)
[[1. 1. 1. ... 1. 1. 1.]
[1. 1. 1. ... 1. 1. 1.]
[1. 1. 1. ... 1. 1. 1.]
...
[1. 1. 1. ... 1. 1. 1.]
[1. 1. 1. ... 1. 1. 1.]
[1. 1. 1. ... 1. 1. 1.]]
import pickle
c1 = pickle.loads(pickle.dumps(c))
print(c1)
c1.large_temporary_array
Container(a=1, b='Ho there', c=2)
---------------------------------------------------------------------------
AttributeError Traceback (most recent call last)
Cell In[15], line 3
1 c1 = pickle.loads(pickle.dumps(c))
2 print(c1)
----> 3 c1.large_temporary_array
AttributeError: 'Container' object has no attribute 'large_temporary_array'
Contexts
The mmfutils.contexts module provides two useful contexts:
NoInterrupt: This can be used to susspend KeyboardInterrupt
exceptions until they can be dealt with at a point that is convenient. A
typical use is when performing a series of calculations in a loop. By
placing the loop in a NoInterrupt context, one can avoid an
interrupt from ruining a calculation:
from mmfutils.contexts import NoInterrupt
complete = False
n = 0
with NoInterrupt() as interrupted:
while not complete and not interrupted:
n += 1
if n > 10:
complete = True
Note: One can nest NoInterrupt contexts so that outer loops are also
interrupted. Another use-case is mapping. See
doc/Animation.ipynb for more examples.
res = NoInterrupt().map(abs, range(-100, 100))
np.sign(res)
array([1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 0, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1,
1, 1])
Interfaces
The interfaces module collects some useful zope.interface tools for checking interface requirements. Interfaces provide a convenient way of communicating to a programmer what needs to be done to used your code. This can then be checked in tests.
from mmfutils.interface import (
Interface,
Attribute,
verifyClass,
verifyObject,
implementer,
)
class IAdder(Interface):
"""Interface for objects that support addition."""
value = Attribute("value", "Current value of object")
# No self here since this is the "user" interface
def add(other):
"""Return self + other."""
Here is a broken implementation. We muck up the arguments to add:
@implementer(IAdder)
class AdderBroken(object):
def add(self, one, another):
# There should only be one argument!
return one + another
try:
verifyClass(IAdder, AdderBroken)
except Exception as e:
print("{0.__class__.__name__}: {0}".format(e))
BrokenMethodImplementation: The object <class '__main__.AdderBroken'> has failed to implement interface __main__.IAdder: The contract of __main__.IAdder.add(other) is violated because 'AdderBroken.add(self, one, another)' requires too many arguments.
Now we get add right, but forget to define value. This is only
caught when we have an object since the attribute is supposed to be
defined in __init__():
@implementer(IAdder)
class AdderBroken(object):
def add(self, other):
return one + other
# The class validates...
verifyClass(IAdder, AdderBroken)
# ... but objects are missing the value Attribute
try:
verifyObject(IAdder, AdderBroken())
except Exception as e:
print("{0.__class__.__name__}: {0}".format(e))
BrokenImplementation: The object <__main__.AdderBroken object at 0x10735a1b0> has failed to implement interface __main__.IAdder: The __main__.IAdder.value attribute was not provided.
Finally, a working instance:
@implementer(IAdder)
class Adder(object):
def __init__(self, value=0):
self.value = value
def add(self, other):
return one + other
verifyClass(IAdder, Adder) and verifyObject(IAdder, Adder())
True
Interface Documentation
We also monkeypatch zope.interface.documentation.asStructuredText()
to provide a mechanism for documentating interfaces in a notebook.
from mmfutils.interface import describe_interface
describe_interface(IAdder)
IAdder
Interface for objects that support addition.
Attributes:
value -- Current value of objectMethods:
add(other) -- Return self + other.
Parallel
The mmfutils.parallel module provides some tools for launching and
connecting to IPython clusters. The parallel.Cluster class
represents and controls a cluster. The cluster is specified by the
profile name, and can be started or stopped from this class:
import logging
logger = logging.getLogger()
logger.setLevel(logging.INFO)
import numpy as np
from mmfutils import parallel
cluster = parallel.Cluster(profile="default", n=3, sleep_time=1.0)
cluster.start()
cluster.wait() # Instance of IPython.parallel.Client
view = cluster.load_balanced_view
x = np.linspace(-6, 6, 100)
y = view.map(lambda x: x**2, x)
print(np.allclose(y, x**2))
cluster.stop()
Waiting for connection file: ~/.ipython/profile_default/security/ipcontroller-client.json
INFO:root:Starting cluster: ipcluster start --daemonize --quiet --profile=default --n=3 2025-10-08 11:45:10.495 [IPController] Hub listening on tcp://127.0.0.1:49245 for registration. 2025-10-08 11:45:10.496 [IPController] Hub using DB backend: DictDB 2025-10-08 11:45:10.752 [IPController] hub::created hub 2025-10-08 11:45:10.754 [IPController] writing connection info to /Users/mforbes/.ipython/profile_default/security/ipcontroller-client.json 2025-10-08 11:45:10.755 [IPController] writing connection info to /Users/mforbes/.ipython/profile_default/security/ipcontroller-engine.json 2025-10-08 11:45:10.756 [IPController] task::using Python leastload Task scheduler 2025-10-08 11:45:10.826 [IPController] Heartmonitor beating every 3000ms 2025-10-08 11:45:11.152 [broadcast-00] BroadcastScheduler 00 started 2025-10-08 11:45:11.152 [broadcast-0] BroadcastScheduler 0 started 2025-10-08 11:45:11.154 [broadcast-01] BroadcastScheduler 01 started 2025-10-08 11:45:11.182 [task] Task scheduler started [leastload] 2025-10-08 11:45:11.183 [IPController] client::client b'x00x80x00Axaa' requested 'connection_request' 2025-10-08 11:45:11.184 [IPController] client::client [b'x00x80x00Axaa'] connected 2025-10-08 11:45:11.187 [IPController] heartbeat::waiting for subscription 2025-10-08 11:45:11.188 [IPController] heartbeat::subscription started Leaving cluster running: /Users/mforbes/.ipython/profile_default/security/cluster-.json INFO:root:waiting for 3 engines INFO:root:0 of 3 running INFO:root:3 of 3 running INFO:root:Stopping cluster: ipcluster stop --profile=default
True
2025-10-08 11:45:18.004 [IPClusterStop] Stopping cluster
2025-10-08 11:45:18.004 [IPClusterStop] Stopping controller
2025-10-08 11:45:18.081 [IPClusterStop] Stopping engine(s): 1759949111
Waiting for connection file: ~/.ipython/profile_default/security/ipcontroller-client.json
If you only need a cluster for a single task, it can be managed with a context. Be sure to wait for the result to be computed before exiting the context and shutting down the cluster!
with parallel.Cluster(profile="default", n=3, sleep_time=1.0) as client:
view = client.load_balanced_view
x = np.linspace(-6, 6, 100)
y = view.map(lambda x: x**2, x, block=True) # Make sure to wait for the result!
print(np.allclose(y, x**2))
Waiting for connection file: ~/.ipython/profile_default/security/ipcontroller-client.json
INFO:root:Starting cluster: ipcluster start --daemonize --quiet --profile=default --n=3 2025-10-08 11:45:39.182 [IPController] Hub listening on tcp://127.0.0.1:49430 for registration. 2025-10-08 11:45:39.183 [IPController] Hub using DB backend: DictDB 2025-10-08 11:45:39.442 [IPController] hub::created hub 2025-10-08 11:45:39.443 [IPController] writing connection info to /Users/mforbes/.ipython/profile_default/security/ipcontroller-client.json 2025-10-08 11:45:39.445 [IPController] writing connection info to /Users/mforbes/.ipython/profile_default/security/ipcontroller-engine.json 2025-10-08 11:45:39.447 [IPController] task::using Python leastload Task scheduler 2025-10-08 11:45:39.490 [IPController] Heartmonitor beating every 3000ms 2025-10-08 11:45:39.842 [broadcast-0] BroadcastScheduler 0 started 2025-10-08 11:45:39.842 [broadcast-00] BroadcastScheduler 00 started 2025-10-08 11:45:39.845 [broadcast-01] BroadcastScheduler 01 started 2025-10-08 11:45:39.868 [task] Task scheduler started [leastload] 2025-10-08 11:45:39.870 [IPController] client::client b'x00x80x00Axaa' requested 'connection_request' 2025-10-08 11:45:39.870 [IPController] client::client [b'x00x80x00Axaa'] connected 2025-10-08 11:45:39.875 [IPController] heartbeat::waiting for subscription 2025-10-08 11:45:39.875 [IPController] heartbeat::subscription started Leaving cluster running: /Users/mforbes/.ipython/profile_default/security/cluster-.json INFO:root:waiting for 3 engines INFO:root:0 of 3 running INFO:root:3 of 3 running INFO:root:Stopping cluster: ipcluster stop --profile=default 2025-10-08 11:45:46.707 [IPClusterStop] Stopping cluster 2025-10-08 11:45:46.707 [IPClusterStop] Stopping controller 2025-10-08 11:45:46.789 [IPClusterStop] Stopping engine(s): 1759949139
Waiting for connection file: ~/.ipython/profile_default/security/ipcontroller-client.json
True
If you just need to connect to a running cluster, you can use
parallel.get_client().
Performance
The mmfutils.performance module provides some tools for high
performance computing. Note: this module requires some additional
packages including
numexp,
pyfftw, and the mkl
package installed by anaconda. Some of these require building system
libraries (i.e. the FFTW). However, the
various components will not be imported by default.
Here is a brief description of the components:
mmfutils.performance.blas: Provides an interface to a few of the scipy BLAS wrappers. Very incomplete (only things I currently need).mmfutils.performance.fft: Provides an interface to the FFTW usingpyfftwif it is available. Also enables the planning cache and setting threads so you can better control your performance.mmfutils.performance.numexpr: Robustly imports numexpr and disabling the VML. (If you don’t do this carefully, it will crash your program so fast you won’t even get a traceback.)mmfutils.performance.threads: Provides some hooks for setting the maximum number of threads in a bunch of places including the MKL, numexpr, and fftw.
Plotting
Several tools are provided in mmfutils.plot:
Fast Filled Contour Plots
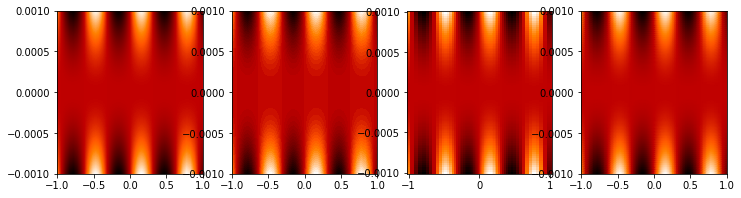
mmfutils.plot.imcontourf is similar to matplotlib’s plt.contourf
function, but uses plt.imshow which is much faster. This is useful
for animations and interactive work. It also supports my idea of saner
array-shape processing (i.e. if x and y have different shapes,
then it will match these to the shape of z). Matplotlib now provides
plt.pcolourmesh which is similar, but has the same interface issues.
%matplotlib inline
from matplotlib import pyplot as plt
import time
import numpy as np
from mmfutils import plot as mmfplt
x = np.linspace(-1, 1, 100)[:, None] ** 3
y = np.linspace(-0.1, 0.1, 200)[None, :] ** 3
z = np.sin(10 * x) * y**2
plt.figure(figsize=(12, 3))
plt.subplot(141)
%time mmfplt.imcontourf(x, y, z, cmap='gist_heat')
plt.subplot(142)
%time plt.contourf(x.ravel(), y.ravel(), z.T, 50, cmap='gist_heat')
plt.subplot(143)
%time plt.pcolor(x.ravel(), y.ravel(), z.T, cmap='gist_heat', shading='auto')
plt.subplot(144)
%time plt.pcolormesh(x.ravel(), y.ravel(), z.T, cmap='gist_heat', shading='gouraud')
CPU times: user 4.97 ms, sys: 1.49 ms, total: 6.46 ms
Wall time: 7.48 ms
CPU times: user 18.2 ms, sys: 2.29 ms, total: 20.5 ms
Wall time: 26.4 ms
CPU times: user 40.6 ms, sys: 1.21 ms, total: 41.8 ms
Wall time: 41.9 ms
CPU times: user 1.34 ms, sys: 70 μs, total: 1.41 ms
Wall time: 1.41 ms
<matplotlib.collections.QuadMesh at 0x1143cc110>
Angular Variables
A couple of tools are provided to visualize angular fields, such as the phase of a complex wavefunction.
%matplotlib inline
from matplotlib import pyplot as plt
import time
import numpy as np
from mmfutils import plot as mmfplt
x = np.linspace(-1, 1, 100)[:, None]
y = np.linspace(-1, 1, 200)[None, :]
z = x + 1j * y
plt.figure(figsize=(9, 2))
ax = plt.subplot(131)
mmfplt.phase_contour(x, y, z, colors="k", linewidths=0.5)
ax.set_aspect(1)
# This is a little slow but allows you to vary the luminosity.
ax = plt.subplot(132)
mmfplt.imcontourf(x, y, mmfplt.colors.color_complex(z))
mmfplt.phase_contour(x, y, z, linewidths=0.5)
ax.set_aspect(1)
# This is faster if you just want to show the phase and allows
# for a colorbar via a registered colormap
ax = plt.subplot(133)
mmfplt.imcontourf(x, y, np.angle(z), cmap="huslp")
ax.set_aspect(1)
plt.colorbar()
mmfplt.phase_contour(x, y, z, linewidths=0.5)
(<matplotlib.contour.QuadContourSet at 0x117505310>,
<matplotlib.contour.QuadContourSet at 0x117530080>)

Debugging
A couple of debugging tools are provided. The most useful is the
debug decorator which will store the local variables of a function
in a dictionary or in your global scope.
from mmfutils.debugging import debug
@debug(locals())
def f(x):
y = x**1.5
z = 2 / x
return z
print(f(2.0), x, y, z)
1.0 2.0 2.8284271247461903 1.0
Mathematics
We include a few mathematical tools here too. In particular, numerical integration and differentiation. Check the API documentation for details.
Developer Instructions
For Developer Notes, please see Notes.md.
Complete code coverage information is provided in
build/_coverage/index.html.
from IPython.display import HTML
ROOTDIR = !hg root
ROOTDIR = ROOTDIR[0]
with open(os.path.join(ROOTDIR, "build/_coverage/index.html")) as f:
coverage = f.read()
HTML(coverage)
No items found using the specified filter.
Change Log
REL: 0.7.5
Add support and testing for Python 3.14.
REL: 0.7.4
Resolve issue #38: Interpolation functions
mmfutils.math.bases.CylindricalBasis.get_PsiandPsiwere incorrectly scaling the wavefunction.
REL: 0.7.3
Removed support for
twist,booset_px, andboost_pxyzinmmfutils.math.bases.Added
factorsto various functions inmmfutils.math.basesallowing independent scaling of each dimension. This will facilitate working with expanding bases. (Resolves issue #37)Fixed an issue with fourier derivatives: We now correctly zero the highest un-paired momentum in states with even numbers of abscissa. (Resolves issue #36)
REL: 0.7.1
Updated
mmfutils.math.bases.interfaces.IBasisKxto haveget_gradientand provided support for this inPeriodicBasisandCylindricalBasis.
REL: 0.7.0
Fully support Python 3.9 through 3.13 with working tests on GitHub CI. (Resolves issue #34.)
Working documentation build on GitLab CI.
Provide a len() method for FPS() so that it works better with e.g. tqdm.
REL: 0.6.7
Add derivative
d=1support for step and mstep. Remove floating point warning.Improved FPS context: better sleeping timing and default timeout behavior.
Drop support for python 3.9 and below. (Could work, but dependencies need careful thought and version pinning.)
Added IBasisCutoff to allow working with the Galerkin projected GPE.
Updated some tests to work with new Numpy formatting (See NEP 51.)
Fixed broken rasterization in contourf but should be unnecessary in the future (see https://github.com/matplotlib/matplotlib/issues/27669).
Improved
performance.auto_timeit.Revert to installing pyfftw from default repo now that issue 303 is fixed.
REL: 0.6.6
Fix issue #31: FFT falbacks should work even if pyfftw is not installed. (Monkeypatch this case in
test_performance_fft.py)Fix issue #32: Make copy of arrays before calling pyfftw builders for the convenience functions to ensure that everything works, even if they are not
WRITEABLE.
REL: 0.6.5
Fix issue #30: measure fft performance and fallback to numpy (with a warning) if it is faster than pyfftw.
REL: 0.6.4
Support python 3.7.13 through 3.11.
Fix some tests.
Add
contexts.FPSwhich is generally preferred toNoInterrupt.Add a
timeout=argument to contexts.Unbind versions.
Fix a couple of bugs in
math.bases.bases.py:Actually use
memoization_GB.PeriodicBasis.kxis now a property.
REL: 0.6.3
Fix some dependencies.
REL: 0.6.2
Fix some issues with GPU and PeriodicBases.
Add warning to FFT plans about data ownership.
Include some Sparkline demonstrations.
Drop support for Python 3.6. (Still no support for 3.10).
REL: 0.6.0
Use Poetry for dependency management.
Update to
src/mmfutilslayout.Better testing and coverage, including GitHub CI.
Odd-numbered lattices are now centered at 0.
Added
fftwextra.
REL: 0.5.4
REL: 0.5.3
Allow Python 3.8. Previous version required python <= 3.7 due to an
issue with
ipyparallel. This
has been resolved with revision 6.2.5 which is available with conda.
REL: 0.5.1
API changes:
Split
mmfutils.containers.ObjectintoObjectBasewhich is simple andObjectMixinwhich provides the picking support. Demonstrate in docs how the pickling can be useful, but slows copying.
REL: 0.5.0
API changes:
Python 3 support only.
mmfutils.math.bases.interfacerenamed tommfutils.math.bases.interfaces.Added default class-variable attribute support to e
mmfutils.containers.Object.Minor enhancements to
mmfutils.math.bases.PeriodicBasisto enhance GPU support.Added
mmfutils.math.bases.interfaces.IBasisLzand support inmmfutils.math.bases.bases.PeriodicBasisfor rotating frames.Cleanup of build environment and tests.
Single environment
_mmfutilsnow used for testing and documentation.
REL: 0.4.13
API changes:
Use
@implementer()class decorator rather thanclassImplementsorimplementsin all interfaces.Improve
NoInterruptcontext. AddedNoInterrupt.unregister(): this allowsNoInterruptto work in a notebook cell even when the signal handlers are reset. (But only works in that one cell.)Added Abel transform
integrate2to Cylindrical bases.
Issues:
Resolved issue #22: Masked arrays work with
imcontourfetc.Resolved issue #23:
NoInterruptworks well except in notebooks due to ipykernel issue #328.Resolved issue #24: Python 3 is now fully supported and tested.
REL: 0.4.10
API changes:
Added
contourf,error_line, andListCollectionstommfutils.plot.Added Python 3 support (still a couple of issues such as
mmfutils.math.integrate.ssum_inline.)Added
mmf.math.bases.IBasisKxand updatelagrangianin bases to acceptk2andkx2for modified dispersion control (along x).Added
math.special.ellipkinv.Added some new
mmfutils.math.linalgtools.
Issues:
Resolved issue #20:
DyadicSumandscipy.optimize.nonlin.JacobianResolved issue #22: imcontourf now respects masked arrays.
Resolved issue #24: Support Python 3.
REL: 0.4.9
< incomplete >
REL: 0.4.7
API changes:
Added
mmfutils.interface.describe_interface()for inserting interfaces into documentation.Added some DVR basis code to
mmfutils.math.bases.Added a diverging colormap and some support in
mmfutils.plot.Added a Wigner Ville distribution computation in
mmfutils.math.wignerAdded
mmfutils.optimize.usolveandubrentqfor finding roots with`uncertanties<https://pythonhosted.org/uncertainties/>`__ support.
Issues:
Resolve issue #8: Use
`ipyparallel<https://github.com/ipython/ipyparallel>`__ now.Resolve issue #9: Use pytest rather than
nose(which is no longer supported).Resolve issue #10: PYFFTW wrappers now support negative
axisandaxesarguments.Address issue #11: Preliminary version of some DVR basis classes.
Resolve issue #12: Added solvers with
`uncertanties<https://pythonhosted.org/uncertainties/>`__ support.




